Bacteria and Bioproduction: Harnessing Resistance Mechanisms for Biotechnology
Introduction
Bacteria are among the most adaptable organisms on Earth, possessing natural mechanisms that allow them to survive and thrive under diverse environmental pressures. Among these mechanisms, resistance systems stand out for their versatility and robustness. While historically studied as survival strategies in microbial ecology, these resistance traits have become invaluable tools in modern biotechnology and synthetic biology. By understanding and repurposing bacterial resistance, scientists are able to create powerful tools for laboratory research, genetic engineering, and industrial bioproduction.
Using Resistance Genes as Genetic Markers
One of the most common applications of bacterial resistance genes is their use as genetic markers. In molecular biology, researchers often link a resistance gene to a target genetic construct, allowing only bacteria that have successfully incorporated the desired DNA to survive under selective conditions. This approach not only streamlines laboratory workflows but also ensures precision and efficiency in experiments involving gene expression, metabolic engineering, or protein production. Resistance genes serve as a reliable indicator, enabling scientists to quickly identify and propagate bacterial strains with specific genetic modifications.
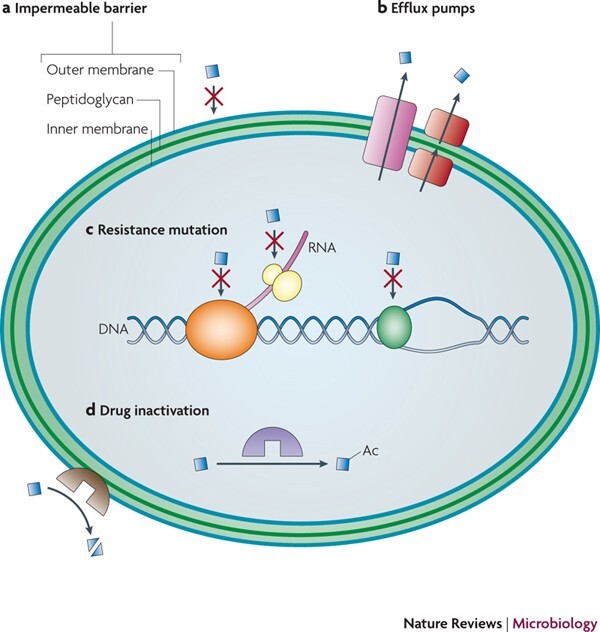
Figure :antibiotic resistance genes in natural environments
Using antibiotic resistance genes as selectable markers revolutionizes bacterial genetic engineering by enabling precise identification of transformed cells. This technique leverages the genes' robust expression to confer survival advantages under antibiotic exposure, streamlining downstream applications in research and biotech.
Common Markers
Aminoglycoside Resistance (e.g., nptII/aph): Encodes neomycin phosphotransferase, inactivating kanamycin and neomycin; widely used in E. coli and plants for plasmid selection due to low environmental transfer risk.
Aminoglycoside Resistance (e.g., aadA): Confers streptomycin/spectinomycin resistance via adenylation; versatile for multi-plasmid systems in Gram-negatives.
Beta-lactam Resistance (e.g., bla): Produces TEM-1 beta-lactamase, hydrolyzing ampicillin; ideal for initial cloning but phased out in GM crops over ecological concerns.
Lab Applications
In cloning, resistance genes on vectors like pUC19 allow ampicillin selection of recombinants post-ligation, achieving >95% positive transformants. Metabolic engineering uses them for pathway optimization, as in biofuel-producing strains where kanR marks high-yield mutants. Protein production workflows (e.g., recombinant insulin in E. coli) rely on dual markers for co-expression, ensuring stable propagation.
Advantages
-
Efficiency: Reduces screening from millions to thousands of colonies via direct plating.
-
Precision: Links survival to successful recombination, minimizing false positives.
-
Versatility: Compatible with CRISPR, transposons, and library construction for functional genomics.
Considerations
While safe for contained labs, genes like nptII pose negligible transfer risks to clinical pathogens per EFSA assessments. Alternatives like auxotrophic markers or fluorescent proteins are gaining traction to avoid resistance dissemination concerns.
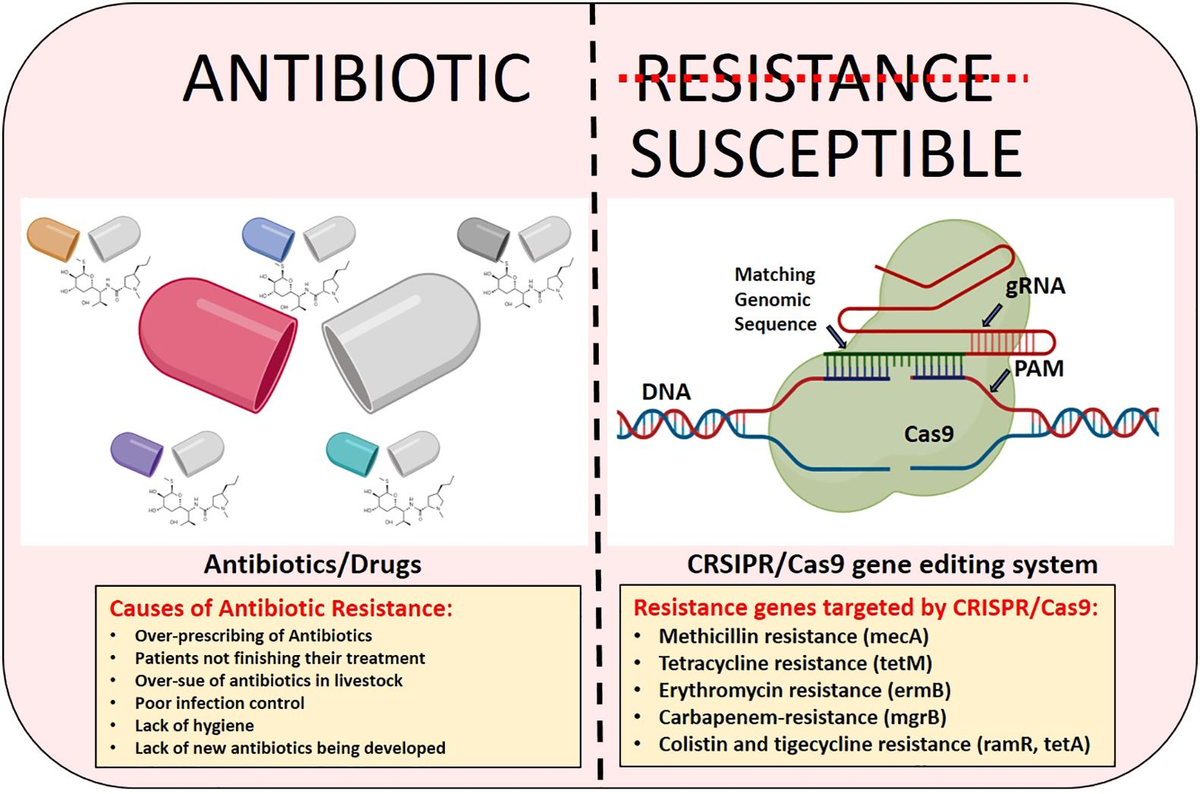
Figure :Illustration of Antibiotic Resistance Mechanisms and CRISPR/Cas9 Gene Editing Applications
Tools for Synthetic Biology and Bacterial Engineering
Antibiotic resistance genes enhance synthetic biology toolkits by integrating into circuits for selection, sensing, and control in engineered bacteria. This expands their utility beyond markers to programmable chassis for industrial bioproducts like biofuels and therapeutics.
Circuit Integration
Resistance modules form kill-switches or biosensors; for example, efflux pumps like tetA respond to inducers, exporting toxins or reporters to monitor environmental cues in metabolic pathways. Paired with CRISPR, they enable precise edits in E. coli or Streptomyces, stabilizing synthetic operons for enzyme cascades in bioplastic monomers (e.g., 1,4-butanediol).
Engineering Applications
In biofuel strains, kanR or cat genes select high-titer butanol producers under dual stress, boosting yields 2-5 fold via directed evolution. Biosensor constructs use bla or aac(3)IV promoters to detect pollutants, triggering metabolite output like indigo for bioremediation. Genome-minimal chassis (e.g., Mycoplasma) incorporate resistance for robustness, supporting quorum-sensing networks that scale production in fermenters.
Key Toolkits
These systems ensure metabolic stability, with orthogonal resistance minimizing off-target effects in cocultures.
>> See this reviewd article
Bioproduction Applications
Resistance mechanisms boost bioproduction by enabling directed evolution of strains with optimized metabolic pathways and stable high-yield genetics. This harnesses bacterial adaptability for scalable manufacturing of enzymes, biofuels, and therapeutics, minimizing development timelines.
Strain Optimization
Adaptive laboratory evolution pairs resistance markers with chemostats, selecting mutants tolerant to toxic intermediates—like isobutanol in biofuel E. coli—yielding 40% higher titers via efflux overexpression or pathway rerouting. Dual-selection systems (e.g., kanR + cat) maintain megaplasmids carrying synthetic operons, preventing segregation during scale-up to 1000L fermenters.
Product Examples
Control Strategies
Biosensors using resistance promoters (e.g., tetA responsive to heavy metals) trigger expression cascades, dynamically allocating flux to products under industrial stress like pH shifts or oxygen limitation. CRISPRi with resistance-coupled guides fine-tunes bottlenecks, as in carotenoid overproduction where silencing competes 5-fold boosts output. These reduce trial-and-error by quantifying fitness via growth rates under selection.
Insights into Microbial Physiology
Studying bacterial resistance in the context of biotechnology provides deeper insights into microbial physiology, gene regulation, and metabolic networks. It allows researchers to explore how bacteria respond to environmental stress, adapt to chemical pressures, and coordinate complex biochemical pathways. These lessons are invaluable for designing synthetic microbial communities, constructing robust industrial strains, and advancing the frontiers of microbial engineering. In essence, bacterial resistance mechanisms, once considered mere survival tools, have become a cornerstone for modern biotechnological applications, demonstrating the extraordinary potential of microorganisms as natural biofactories.
Conclusion
Harnessing these natural systems bridges the gap between fundamental microbiology and applied biotechnology. It showcases how evolutionary adaptations, shaped over millions of years, can be transformed into tools for human innovation, enabling breakthroughs in research, industrial bioproduction, and synthetic biology. Bacteria, with their intricate resistance strategies, continue to inspire new ways of thinking about molecular engineering, genetic design, and the production of bioactive molecules, solidifying their role as indispensable partners in biotechnology and laboratory science.
Recent Posts
-
Advances in Peptide Synthesis Technologies and Their Applications in Modern Biotechnology
Advances in Peptide Synthesis Technologies and Their Applications in Modern Biotechnology …27th Mar 2026 -
Western Blot
Introduction Western blotting is a fundamental analytical technique widely used in molecular biology …27th Mar 2026 -
Ethidium Bromide as a Cooperative Effector of DNA Structure
Introduction DNA structure is not static. Under specific physicochemical conditions, DNA molecules c …25th Mar 2026