Choosing the Right CRISPR/Cas9 Kit for Your Experiment: A Complete Guide
Choosing the Right CRISPR/Cas9 Kit for Your Experiment: A Complete Guide
Introduction
Genome editing has undergone a profound transformation with the emergence of CRISPR/Cas9 technology, enabling researchers to manipulate genetic material with remarkable precision and efficiency. However, the rapid expansion of available CRISPR kits and genome editing tools has introduced a new challenge: selecting the most appropriate system for a specific experimental context. Making the right choice is not simply a matter of convenienceit is a critical factor that determines the success, reproducibility, and accuracy of your results.
Understanding the Basics of CRISPR/Cas9 Systems
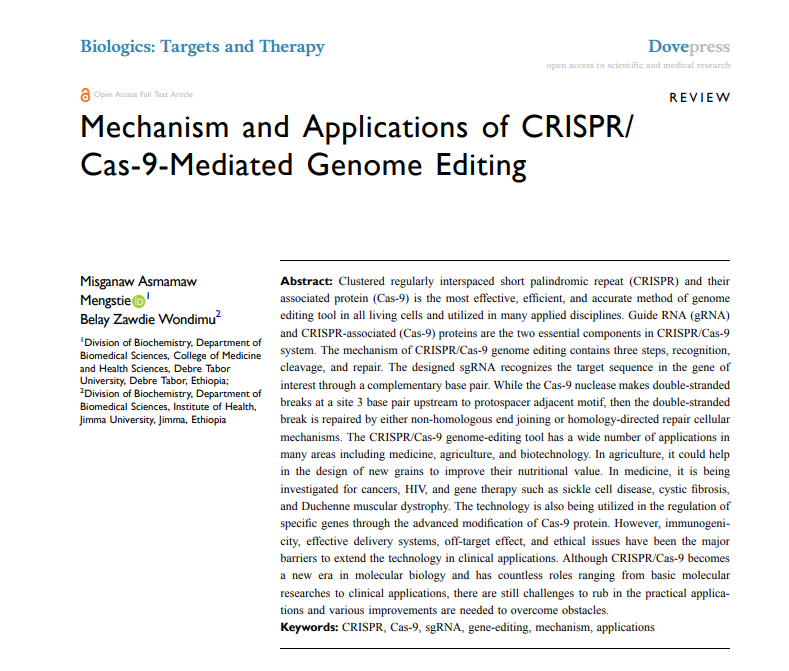
At its core, CRISPR/Cas9 technology relies on a highly coordinated molecular mechanism involving the Cas9 nuclease and a guide RNA (gRNA). The gRNA directs the Cas9 enzyme to a precise genomic location, where it introduces a double-strand break in the DNA. This break subsequently activates the cell’s natural repair pathways, primarily non-homologous end joining (NHEJ) or homology-directed repair (HDR). While NHEJ often leads to gene disruption through insertions or deletions, HDR enables precise sequence modifications when a donor template is present. The efficiency and outcome of these processes are strongly influenced by the design and quality of the CRISPR kit used, making it essential to understand these underlying mechanisms before making a selection.
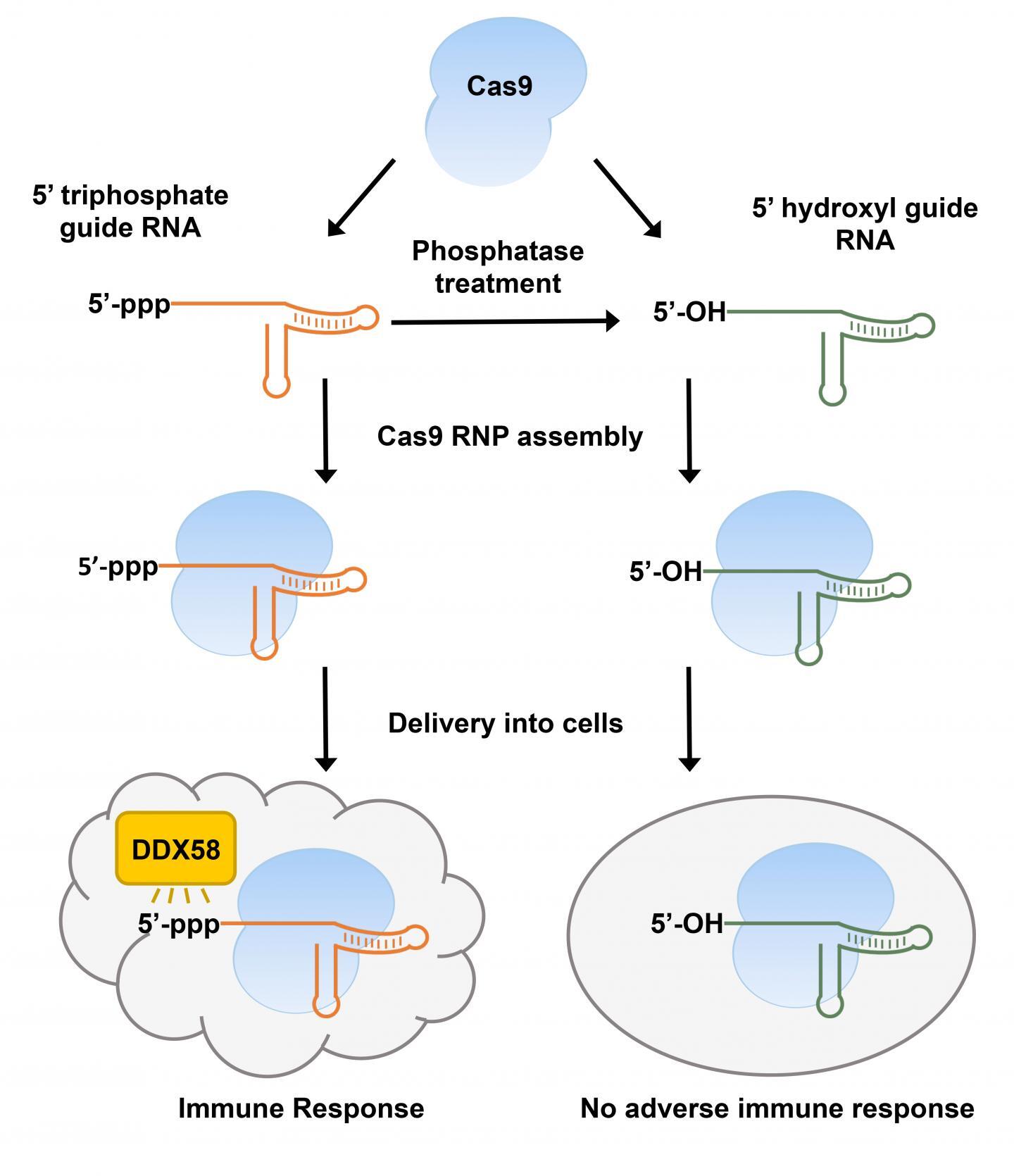
Figure: In vitro transcribed-CRISPR guide RNAs trigger innate immune response in cells, but can be prevented by removing the triphosphate moiety
Defining Your Experimental Objective
A clear definition of your experimental goal is the foundation of any successful CRISPR experiment. Whether the objective is to generate a gene knockout, introduce a precise mutation, regulate gene expression, or perform large-scale functional screening, each application requires a specific configuration of CRISPR components. For instance, simple knockout experiments may rely on standard Cas9 systems exploiting NHEJ, whereas precise genome editing demands HDR-compatible kits along with carefully designed donor templates. Similarly, gene regulation studies require modified systems such as CRISPR interference or activation platforms. Without aligning the kit selection to the experimental purpose, even the most advanced tools may fail to deliver meaningful results.
Choosing the Right Delivery Method
The method used to deliver CRISPR components into target cells plays a decisive role in determining editing efficiency and specificity. Plasmid-based systems, which encode both Cas9 and gRNA, remain widely used due to their simplicity and accessibility, making them suitable for routine applications. However, their reliance on intracellular expression can introduce delays and increase the risk of off-target activity. In contrast, ribonucleoprotein (RNP) complexes, which consist of pre-assembled Cas9 protein and gRNA, offer immediate functionality upon delivery and significantly reduce unintended modifications. Viral delivery systems, including lentiviral and adeno-associated virus vectors, provide robust solutions for difficult-to-transfect cells or in vivo studies, although they require more complex handling and careful consideration of biosafety. The choice of delivery method must therefore balance efficiency, safety, and experimental constraints.
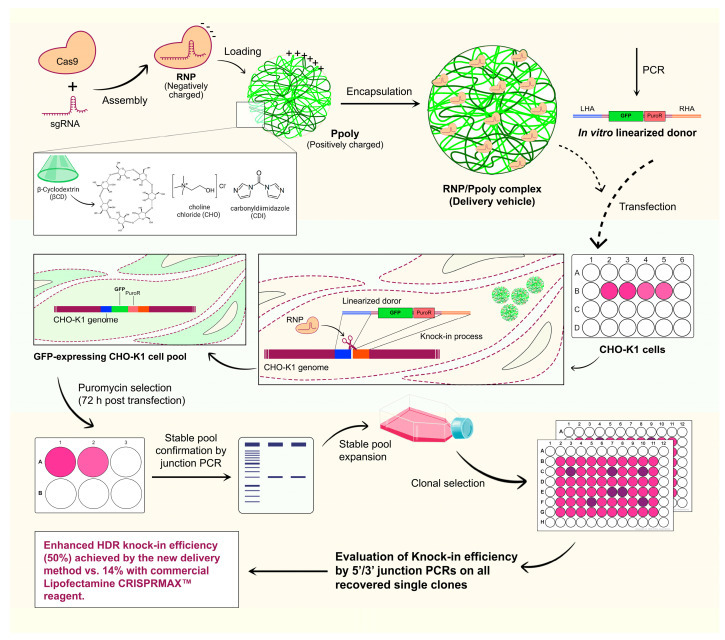
Figure: Schematic Overview of the Targeted Integration with Linearized dsDNA (TILD)-CRISPR Knock-In (KI) Strategy Using the Cationic Hyper-Branched Cyclodextrin-Based Polymer (Ppoly).
Matching the Kit to Your Cell Type
The biological model used in your experiment is another crucial determinant in selecting an appropriate CRISPR kit. While immortalized cell lines are generally permissive and compatible with a wide range of delivery systems,
primary cells and stem cells present additional challenges due to their sensitivity and lower transfection efficiencies. In such cases, high-performance delivery methods like RNP complexes or optimized viral systems are often
required to achieve satisfactory editing outcomes. For in vivo applications, delivery becomes even more complex, necessitating the use of specialized vectors or nanoparticle-based approaches. A mismatch between the CRISPR system and the target cell type can significantly compromise experimental success, highlighting the importance of careful consideration at this stage. 
Ensuring High Editing Efficiency and Specificity
Achieving high editing efficiency while minimizing off-target effects remains one of the central challenges in genome editing. Advances in CRISPR technology have led to the development of high-fidelity Cas9 variants and improved guide RNA design tools that enhance targeting precision. Many modern kits incorporate pre-validated components and optimized protocols to ensure consistent performance across different experimental conditions. Nevertheless, researchers must remain vigilant, as even minor inefficiencies or unintended edits can impact downstream analyses. Selecting a kit that prioritizes both efficiency and specificity is therefore essential for generating reliable and reproducible data.
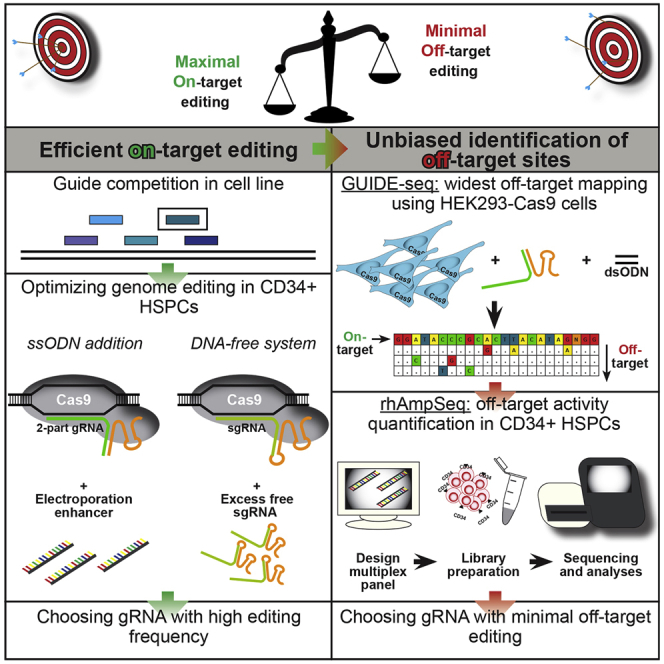
Figure: Optimizing CRISPR-Cas9 for Safe and Efficient Genome Editing in Hematopoietic Stem Cells
Workflow Simplicity and Technical Support
Beyond technical performance, the overall usability of a CRISPR kit can significantly influence the experimental workflow. Ready-to-use kits with pre-designed components and comprehensive protocols can greatly reduce setup
time and minimize variability, particularly for laboratories with limited experience in genome editing. In addition, access to reliable technical support and detailed documentation can facilitate troubleshooting and improve overall success rates. While more customizable systems may offer greater flexibility, they often require a higher level of expertise, making it important to align the complexity of the kit with the capabilities of the research team.![]()
Scaling Your Experiment
The scale of your experiment is another factor that should guide your choice of CRISPR tools. Small-scale studies focused on individual genes can typically be conducted using standard kits, whereas large-scale functional genomics
projects require more sophisticated solutions such as pooled CRISPR libraries. These high-throughput systems are designed to enable the simultaneous targeting of thousands of genes, often in combination with automated workflows and advanced data analysis pipelines. Selecting a kit that aligns with the scale of your research ensures both efficiency and cost-effectiveness.![]()
Common Pitfalls to Avoid
Despite its versatility, CRISPR technology is not without challenges, and several common pitfalls can undermine experimental success. One of the most frequent mistakes is selecting a kit without considering compatibility with the target cell type or experimental objective. Additionally, insufficient attention to guide RNA design and off-target effects can lead to inaccurate or misleading results. The use of poorly validated reagents further compounds these issues, emphasizing the importance of choosing high-quality kits from reputable sources. Careful planning and a thorough understanding of the system are essential to avoid these common errors.
Validation: A Critical Step
Validation is a fundamental component of any genome editing experiment, ensuring that the intended modifications have been successfully introduced. Molecular techniques such as PCR and DNA sequencing are commonly used to confirm genetic changes, while protein-level analyses, including Western blotting or ELISA, provide insight into functional outcomes. In many cases, additional phenotypic or functional assays are required to fully assess the biological impact of the edits. Without rigorous validation, it is impossible to confidently interpret experimental results, making this step indispensable in the CRISPR workflow.
Future Perspectives in Genome Editing
The field of genome editing continues to evolve rapidly, with new technologies expanding the capabilities of CRISPR-based systems. Innovations such as base editing and prime editing allow for precise nucleotide changes without introducing double-strand breaks, reducing the risk of unintended effects. Additionally, epigenome editing approaches enable the modulation of gene expression without altering the underlying DNA sequence. As these tools become more widely available, they are expected to further enhance the precision and versatility of genome engineering, opening new avenues for research and therapeutic applications.
Recent advances have further expanded the capabilities of the CRISPR-Cas system. This article highlights its latest applications, explores future perspectives, and addresses the limitations that still challenge researchers in genome editing :
Conclusion
Selecting the right CRISPR/Cas9 kit is a strategic decision that extends far beyond simple product choice. It requires a comprehensive understanding of the experimental objective, biological system, and technical constraints involved. By carefully evaluating these factors and choosing a kit that balances efficiency, specificity, and usability, researchers can significantly improve the likelihood of successful genome editing. As the technology continues to advance, informed decision-making will remain a key driver of innovation and scientific discovery.
Recent Posts
-
Western Blot in Single-Cell Proteomics: Unlocking Protein Insights One Cell at a Time
Western Blot in Single-Cell Proteomics: Unlocking Protein Insights One Cell at a Time Proteomics has …17th Mar 2026 -
Ethical Considerations in Genome Editing Technologies
Ethical Considerations in Genome Editing Technologies Genome editing technologies, particularly CRIS …17th Mar 2026 -
Top Considerations When Buying Genome Editing Reagents
Top Considerations When Buying Genome Editing Reagents The success of any genome editing experiment …17th Mar 2026