Ethical Considerations in Genome Editing Technologies
Ethical Considerations in Genome Editing Technologies
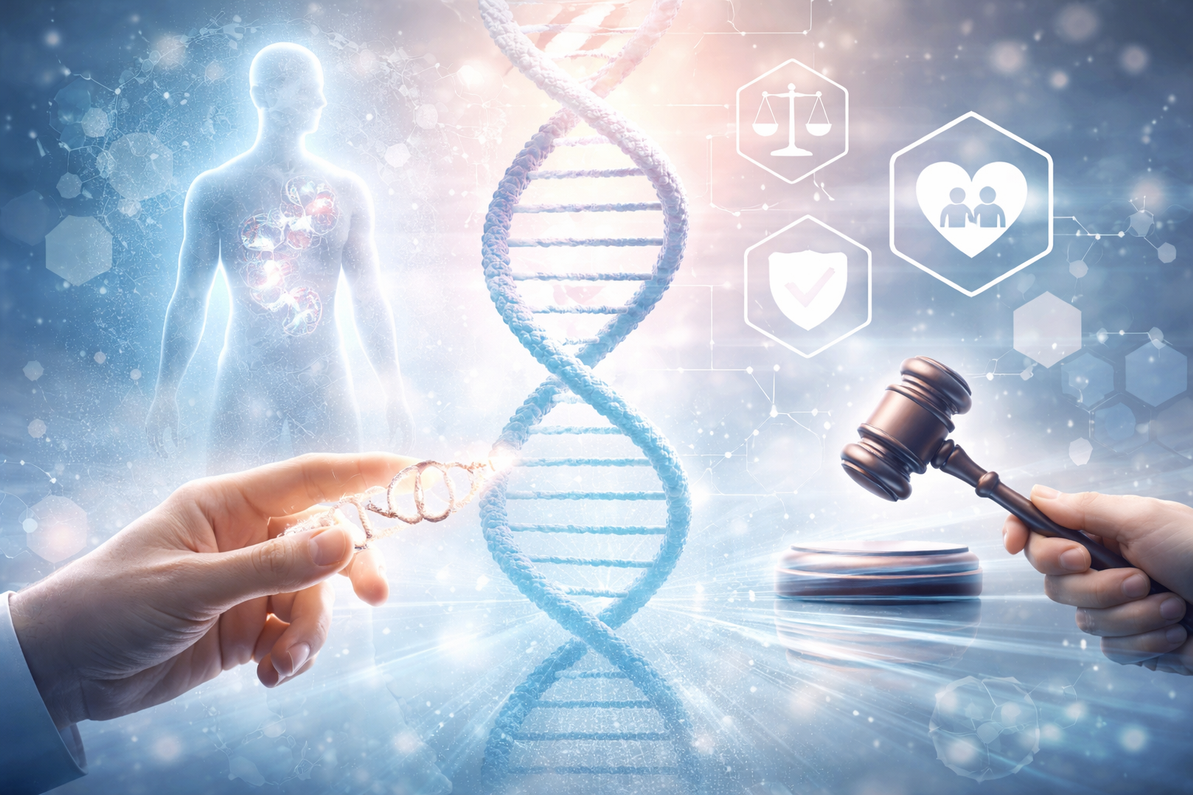
Genome editing technologies, particularly CRISPR/Cas9, have revolutionized modern biology, providing researchers with unprecedented control over the genetic code. From correcting disease-causing mutations to engineering crops with enhanced resilience, the possibilities appear limitless. Yet, alongside these extraordinary scientific advancements arise profound ethical questions that cannot be overlooked. The power to rewrite life itself comes with responsibilities, challenges, and societal implications that require careful consideration.
Balancing Innovation and Responsibility
At the heart of the ethical debate is the tension between innovation and responsibility. Genome editing holds tremendous promise for alleviating human suffering through therapies for genetic diseases, cancer, and rare disorders. However, the same tools that can correct a harmful mutation could also be used to modify non-disease traits, raising concerns about "designer babies," genetic inequality, or unforeseen consequences on future generations. Researchers and institutions must weigh the potential benefits against possible harms, ensuring that experimentation proceeds with caution, transparency, and strict oversight.
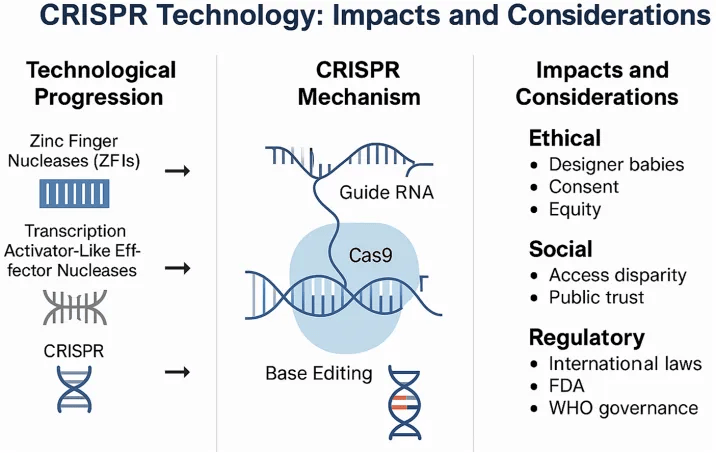
➙ CRISPR-Cas9 revolutionized the field by offering a more efficient, programmable, and accessible approach to genome editing
Safety and Unintended Consequences
One of the central ethical challenges in genome editing is safety. Even the most precise technologies carry risks of off-target effects, unintended mutations, or long-term impacts that are difficult to predict. Editing the human germline, which passes changes to offspring, introduces a layer of complexity and moral responsibility that extends far beyond the individual patient. Ethical decision-making in genome editing requires robust preclinical studies, careful risk assessment, and ongoing monitoring to minimize potential harm, while acknowledging that absolute certainty is impossible in any experimental setting.
➙ The table summarizes current computational approaches for predicting off-target effects in CRISPR/Cas9 gene editing. As highlighted in the article Off-target effects in CRISPR/Cas9 gene editing, these prediction algorithms mainly rely on sgRNA sequences, which can bias their outputs toward sgRNA-dependent off-targets. Moreover, most in silico tools do not fully account for the complex intranuclear environment, including epigenetic marks and chromatin organization, meaning that experimental validation is still essential to confirm predicted off-target sites.
| Methods | Characteristics | Advantages | Disadvantages | ||
|---|---|---|---|---|---|
| In silico prediction | Alignment based models | CasOT [21] | Adjustable in PAM sequence and the mismatch number (at most 6) | Conveniently accessable via internet | Biased toward sgRNA-dependent off-target effects; results need experimental validation |
| Cas-OFFinder [22] | Adjustable in sgRNA length, PAM type, and number of mismatches or bulges | ||||
| FlashFry [23] | Provides information about GC contents | ||||
| Crisflash [24] | High in speed | ||||
| Scoring based models | MIT [15, 25] | Based on the position of the mismatches to the gRNA | |||
| CCTop [26] | Based on the distances of the mismatches to the PAM | ||||
| CROP-IT [27] | |||||
| CFD [28] | Based on a experimentally validated dataset | ||||
| DeepCRISPR [29] | Considers both sequence and epigenetic feature | ||||
| Elevation [30] | |||||
| Experimental detection | Cell-free methods | Digenome-seq [31–33] | Digests purified DNA with Cas9/gRNA RNP → WGS | Highly sensitive | Expensive; requires high sequencing coverage; requires a reference genome |
| DIG-seq [34] | Uses cell-free chromatin with Digenome-seq pipeline | Concerning chromatin accessibility; higher validation rate than Digenome-seq | |||
| Extru-seq [35] | Pre-incubates live cells with Cas9/sgRNA RNP complex→rapidly kill cells by extruder→WGS | Low miss rate; low false positive rate | Expensive; difficult to detect Cas9-mediated large deletions, chromosomal depletions, and translocations | ||
| SITE-seq [37] | A biochemical method with selective biotinylation and enrichment of fragments after Cas9/gRNA digestion | Minimal read depth; eliminated background; does not require a reference genome | Low sensitivity; low validation rate | ||
| CIRCLE-seq [38–40] | Circularizes sheared genomic DNA→incubate with Cas9/gRNA RNP→linearized DNA for NGS | ||||
| Cell culture-based methods | WGS [41–43] | Sequences the whole genome before and after gene editing | Comprehensive analysis of the whole genome | Expensive; limited number of clones can be analyzed | |
| ChIP-seq [44–47] | Analyzes binding sites of catalytically inactive dCas9 | Detection of Cas9 binding sites genome-wide | Low validation rate; affected by antibody specificity and chromatin accessibility | ||
| IDLV [48–52] | Integrates IDLV into DSBs | Detects off-targets in cells that are difficult to transfect | Low sensitivity; high false positive rate | ||
| GUIDE-seq [36, 53–55] | Integrates dsODNs into DSBs | Highly sensitive, low in cost, low false positive rate | Limited by transfection efficiency | ||
| LAM–HTGTS [57–59] | Detects DSB-caused chromosomal translocations by sequencing bait-prey DSB junctions | Accurately detects chromosomal translocations induced by DSBs | Only detects DSBs with translocation; efficiency limited by chromatin accessibility | ||
| BLESS [60, 61] | Captures DSBs in situ by biotinylated adaptors | Directly capture DSBs in situ | Only identifies off-target sites at the time of detection | ||
| BLISS [61, 62] | Captures DSBs in situ by dsODNs with T7 promoter sequence | Directly capture DSBs in situ; low-input needed | |||
| In vivo detection | Discover-seq [63] | Utilizes DNA repair protein MRE11 as bait to perform ChIP-seq | Highly sensitive; high precision in cells | Has false positives | |
| GUIDE-tag [64] | Uses biotin-dsDNA to mark DSBs | Highly sensitive; detects off target sites in vivo | The incorporation rate of biotin-dsDNA is relatively low (∼6%) |
Equity and Accessibility
Ethical considerations in genome editing also include questions of equity and accessibility. As therapies and technologies advance, there is a risk that only wealthy individuals or nations will benefit, exacerbating existing disparities in healthcare and social opportunity. Should powerful genome editing technologies be restricted to those who can afford them, or should there be policies to ensure broader access? Addressing these questions is essential to prevent widening societal inequalities and to ensure that the benefits of scientific progress are shared responsibly.
Informed Consent and Autonomy
Respecting autonomy and securing informed consent are fundamental ethical principles in any biomedical intervention. Genome editing research, particularly involving human subjects, must guarantee that participants fully understand the scope, risks, and potential long-term consequences of the procedures. In the context of germline editing, where changes may affect future generations, informed consent becomes more complex, raising questions about who can consent on behalf of those yet unborn and how society should approach this challenge.
Regulatory Oversight and Governance
The rapid pace of genome editing research has outstripped the development of regulatory frameworks in many regions. Effective governance is essential to balance scientific freedom with ethical responsibility. International guidelines, institutional review boards, and government regulations play crucial roles in setting standards for safety, ethical conduct, and transparency. Global collaboration and harmonization of rules can help prevent unethical experimentation while fostering responsible innovation.
Societal Implications and Public Engagement
Genome editing is not only a scientific issue but also a societal one. Public perception, cultural values, and ethical norms vary widely, influencing how technologies are received and regulated. Engaging the public through education, dialogue, and participatory decision-making ensures that genome editing progresses in a way that reflects societal priorities and values. Scientists have a responsibility to communicate the potential benefits and risks of genome editing in accessible terms, fostering informed debate rather than secrecy or hype.
Striking a Balance
Ultimately, the ethical considerations surrounding genome editing demand a delicate balance. Researchers must pursue innovation and discovery while upholding principles of safety, equity, transparency, and respect for human dignity. Ethical reflection is not a barrier to progress; rather, it ensures that scientific advancements contribute positively to society and minimize harm. By integrating ethical deliberation into research planning, policy development, and public engagement, genome editing can fulfill its transformative potential responsibly.
Fig. 1: Limitations of short-read sequencing in gene editing analysis.
Conclusion
Genome editing technologies offer unparalleled opportunities to improve human health, agriculture, and scientific understanding. Yet, with great power comes great responsibility. Ethical considerations ranging from safety and consent to equity and societal impact must guide the development and application of these tools. Thoughtful, proactive engagement with these issues ensures that genome editing remains not only a scientific achievement but also a moral one, capable of advancing society without compromising our shared values.
Recent Posts
-
Western Blot in Single-Cell Proteomics: Unlocking Protein Insights One Cell at a Time
Western Blot in Single-Cell Proteomics: Unlocking Protein Insights One Cell at a Time Proteomics has …17th Mar 2026 -
Ethical Considerations in Genome Editing Technologies
Ethical Considerations in Genome Editing Technologies Genome editing technologies, particularly CRIS …17th Mar 2026 -
Top Considerations When Buying Genome Editing Reagents
Top Considerations When Buying Genome Editing Reagents The success of any genome editing experiment …17th Mar 2026