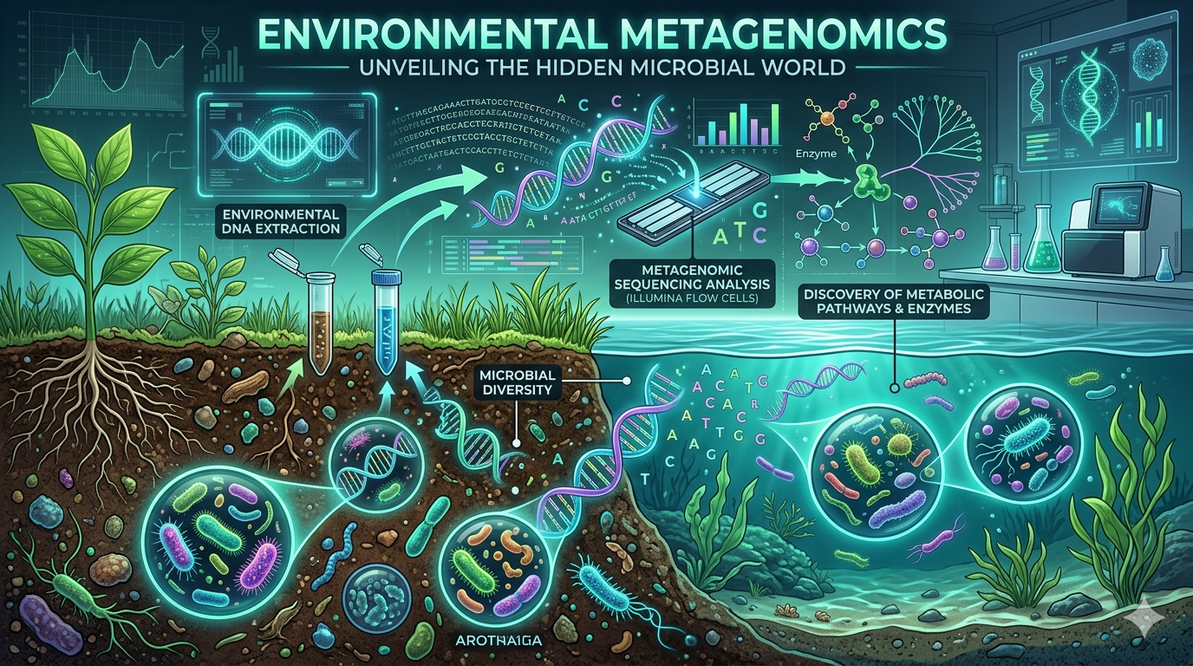
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds
Introduction
Microorganisms are the unseen architects of Earth's ecosystems. From the deepest ocean trenches to nutrient-rich soils, microbial communities play a critical role in biogeochemical cycles, environmental stability, and ecosystem productivity. Traditional microbiology techniques, relying on culturing individual species in the laboratory, have long provided insights into bacterial physiology and genetics. However, the vast majority of microbes estimated at over 99% cannot be easily cultivated under standard laboratory conditions. This limitation has historically constrained our understanding of microbial diversity and function in natural environments. Metagenomics has revolutionized this field, allowing scientists to study entire microbial communities directly from environmental samples without the need for cultivation.
What is Metagenomics?
Metagenomics refers to the direct analysis of genetic material recovered from environmental samples. Instead of isolating individual species, metagenomics captures all DNA present in a community, providing a comprehensive snapshot of the microbial population. This approach enables researchers to identify species composition, functional genes, metabolic pathways, and even novel enzymes or bioactive compounds. By bypassing the cultivation step, metagenomics reveals the full complexity of microbial ecosystems and offers a deeper understanding of microbial diversity, evolution, and ecological interactions.
Environmental microbiology leverages metagenomics to study diverse habitats such as soil, freshwater, marine ecosystems, sediments, and extreme environments. By sequencing environmental DNA, scientists can detect rare or previously unknown species, uncover functional genes, and map interactions between microorganisms that were previously hidden from view.
Methods in Metagenomics
Metagenomics employs optimized DNA extraction, high-throughput sequencing, and bioinformatics for microbial community analysis. Key methods include amplicon sequencing for taxonomy and shotgun sequencing for functional insights. These techniques reveal novel genes and pathways with biotech potential.
Core Workflow
DNA extraction from environmental samples targets diverse microbes like bacteria, archaea, fungi, and viruses, often using mechanical or enzymatic pretreatments for high yield and purity. High-throughput platforms such as Illumina (short reads, low error) or Oxford Nanopore (long reads) generate millions of fragments. Bioinformatics pipelines handle assembly, gene annotation, and functional prediction, with depth influencing genome recovery.
Amplicon Sequencing
This targets marker genes like 16S rRNA (bacteria/archaea) or ITS/18S rRNA (fungi) to profile species composition and abundance. It offers cost-effective diversity assessment but struggles with species-level resolution. Commonly used in early studies for community snapshots.
Shotgun Metagenomics
Random sequencing of all sample DNA provides taxonomic profiles plus functional data on enzymes, resistance genes, and pathways. Enhancements like Hi-C or single-cell approaches improve assembly from complex samples. Superior for discovering novel biotech elements over amplicon methods.
Emerging Advances
Long-read sequencing demands high-molecular-weight DNA, with customized extraction protocols boosting accuracy. Hybrid approaches combine methods for better resolution in microbiome research. Applications span environmental monitoring, antibiotic discovery, and human gut studies.
Applications in Environmental Microbiology
Metagenomics has transformed environmental microbiology in several key areas:
-
Discovery of New Enzymes and Biocatalysts
Environmental metagenomes are a rich source of enzymes with unique properties, including thermostable, solvent-tolerant, or high-activity enzymes. These enzymes have applications in industrial biotechnology, biofuel production, and green chemistry. For instance, cellulases, lipases, and proteases identified through metagenomic studies are now used in processes ranging from waste biomass degradation to pharmaceutical manufacturing. -
Mapping Metabolic Pathways
By analyzing functional genes within microbial communities, researchers can reconstruct metabolic networks that drive nutrient cycling, carbon fixation, nitrogen metabolism, and other essential ecosystem functions. These insights are critical for understanding soil fertility, biogeochemical cycles, and the role of microbes in environmental sustainability. -
Resistome Studies and Environmental Health
Metagenomics allows for the detection of resistance genes in environmental bacteria, collectively known as the environmental resistome. This knowledge is crucial for tracking the spread of antimicrobial resistance, understanding microbial adaptation, and developing strategies to mitigate resistance gene propagation in natural and agricultural ecosystems. -
Microbial Ecology and Community Dynamics
Metagenomics provides a window into how microbial communities interact with each other and their environment. By comparing metagenomic profiles across samples and conditions, scientists can study community succession, resilience to stress, and responses to environmental change. This has implications for climate change studies, pollution remediation, and ecosystem management. -
Bioremediation and Environmental Biotechnology
Functional metagenomics can identify microbes capable of degrading pollutants, such as hydrocarbons, pesticides, or heavy metals. This allows for the design of microbial consortia for bioremediation and environmental cleanup, turning natural microbial diversity into practical solutions for human challenges.
Challenges and Future Directions
While metagenomics has opened unprecedented opportunities, it also presents challenges. Environmental samples are complex, containing DNA from thousands of species, which makes assembly and annotation difficult. Short-read sequencing may fragment genomes, complicating the reconstruction of complete microbial pathways. Additionally, interpreting functional potential from sequence data requires careful experimental validation.
Future developments in long-read sequencing, single-cell genomics, and improved bioinformatics tools are expected to overcome these limitations. Integrating metagenomics with metatranscriptomics, metabolomics, and proteomics will provide a more complete picture of microbial function and activity. This multi-omics approach will enhance our ability to harness microbial communities for biotechnological applications, environmental monitoring, and sustainable industrial processes.
Conclusion
Metagenomics has fundamentally transformed environmental microbiology by providing access to the full genetic diversity of microbial communities without cultivation. From the discovery of novel enzymes to mapping metabolic networks and studying the environmental resistome, metagenomics enables scientists to explore hidden microbial worlds with unprecedented depth. As sequencing technologies and computational methods continue to advance, metagenomics will remain a cornerstone of environmental microbiology, unlocking new opportunities in biotechnology, ecosystem management, and microbial ecology. By understanding and leveraging microbial diversity, we can harness nature’s genetic potential to drive innovation and sustainability in science and industry.
Recent Posts
-
The Intestinal Microbiota: A Complex Microbial Ecosystem
The Intestinal Microbiota: A Complex Microbial Ecosystem The intestinal microbiota represents one of …13th Mar 2026 -
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds Introduction Microorga …13th Mar 2026 -
The Environmental Resistome: Hidden Genetic Libraries in Soil and Water Microbiomes
The Environmental Resistome : Hidden Genetic Libraries in Soil and Water Microbiomes Introduction Ba …13th Mar 2026