The Environmental Resistome: Hidden Genetic Libraries in Soil and Water Microbiomes
The Environmental Resistome : Hidden Genetic Libraries in Soil and Water Microbiomes
Introduction
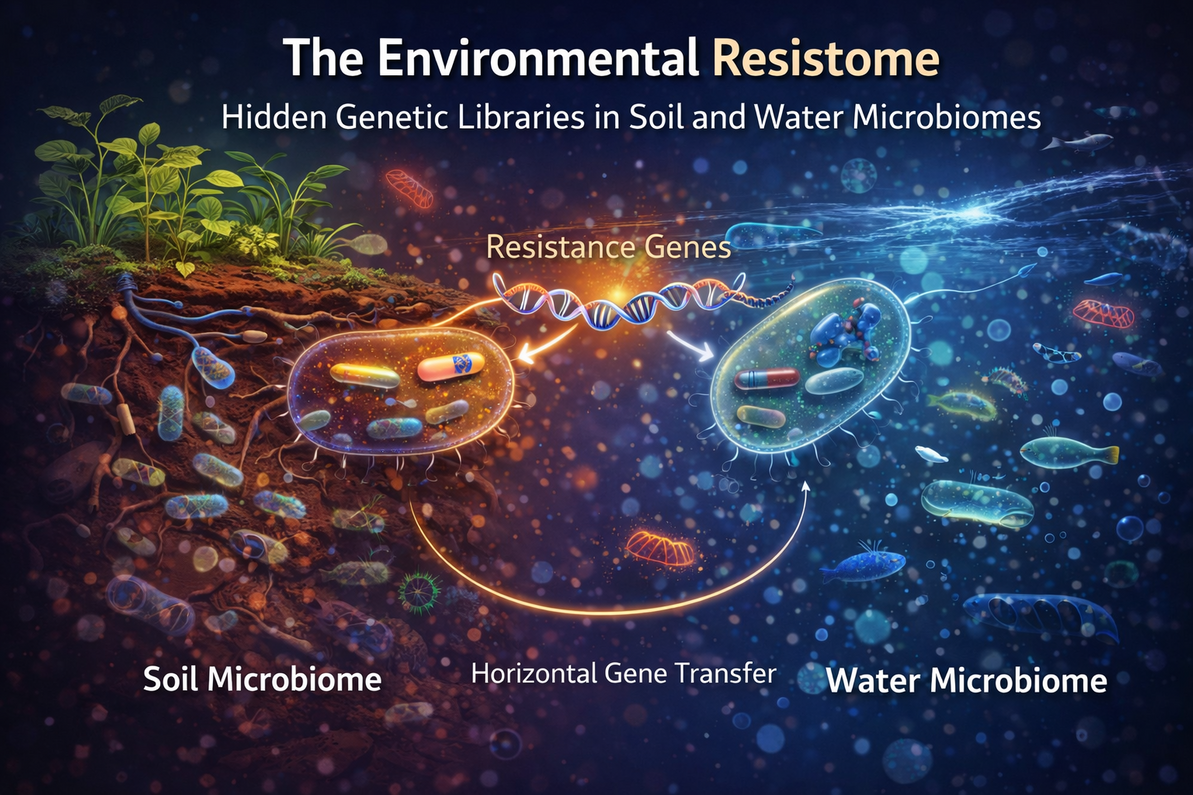
Bacteria in soil and aquatic ecosystems are not just passive inhabitants they are dynamic reservoirs of genetic information, including genes that confer resistance to various chemical compounds. This collection of resistance genes is often referred to as the environmental resistome, representing an extensive natural library of adaptive traits that bacteria can exchange, evolve, and use to survive in competitive ecosystems. Understanding the environmental resistome is critical for microbiologists and biotechnologists, as it provides insight into microbial evolution, genetic diversity, and potential biotechnological applications.
What is the Environmental Resistome?
The environmental resistome refers to the full collection of antibiotic resistance genes (ARGs) and related genetic elements found in microbial communities across natural and human-influenced environments, such as soil, water bodies, sediments, and air.
Core Definition
These genes encode mechanisms that help bacteria resist antimicrobials, including naturally produced antibiotics, heavy metals, and other toxins, predating clinical antibiotic use by millennia. Unlike the clinical resistome, which focuses on pathogens in humans or animals, the environmental resistome acts as a vast, ancient reservoir shaped by ecological pressures and horizontal gene transfer (HGT) via plasmids, transposons, and other mobile elements.
Key Locations
It is ubiquitous in diverse habitats.
Soil: A primary hotspot due to diverse bacteria and natural antibiotic producers.
Aquatic systems: Rivers, lakes, oceans, and wastewater harbor ARGs, enriched by runoff.
Other niches: Air, sediments, and built environments also contribute.
Significance
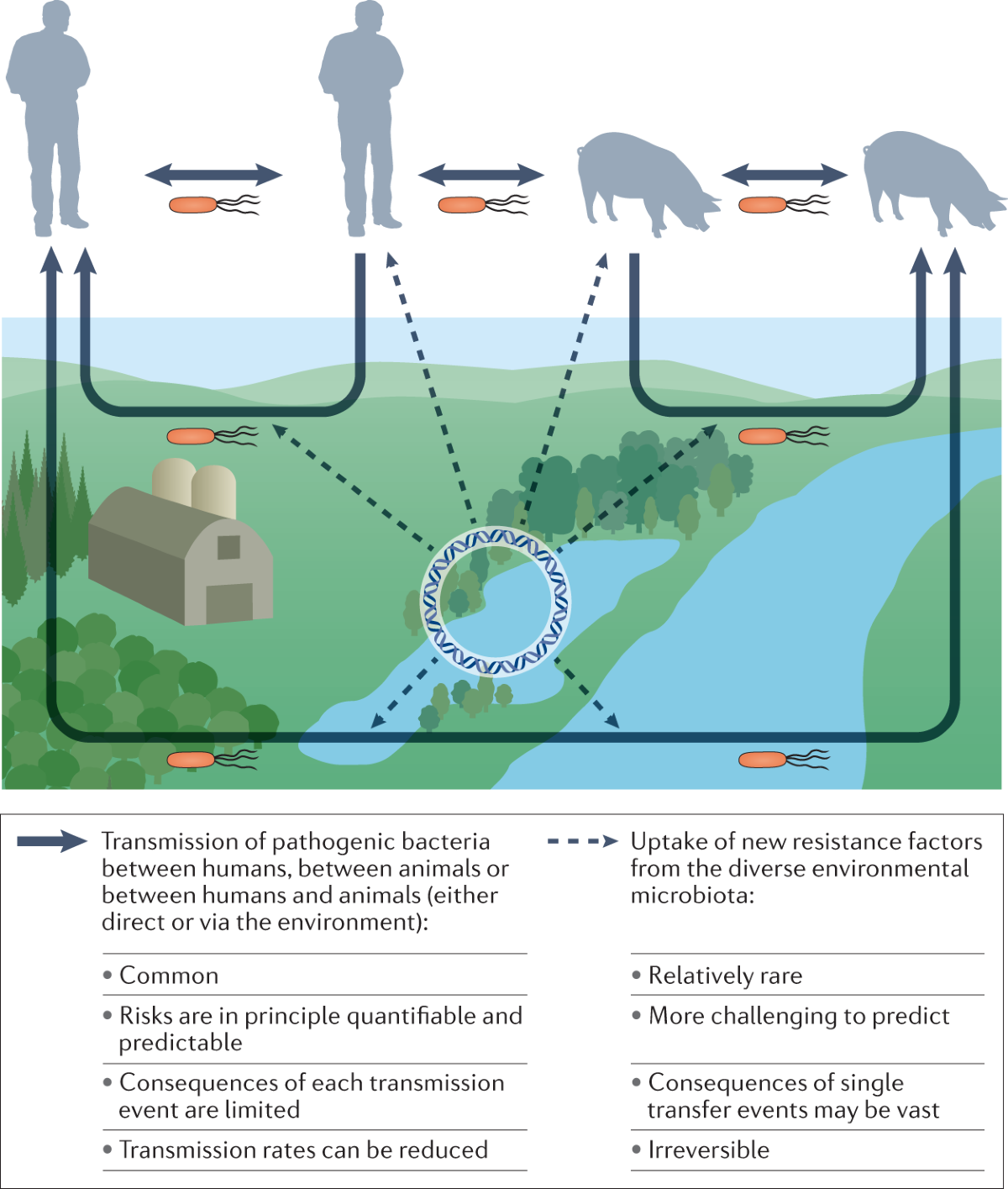
The resistome contributes to bacterial evolution and can transfer to clinical pathogens via HGT mechanisms like transformation, conjugation, and transduction, fueling antimicrobial resistance (AMR) under the One Health framework. Human activities, such as agriculture and wastewater discharge, amplify its spread, posing risks to health, food security, and ecosystems. Recent metagenomic studies highlight its global connectivity across habitats.
Read more >>
Soil Microbiomes as Genetic Reservoirs
Soil microbiomes serve as major genetic reservoirs for antibiotic resistance genes (ARGs), driven by their immense microbial diversity and density, which foster evolutionary pressures from natural antimicrobials.
Microbial Drivers
Genera like Streptomyces, Bacillus, and Pseudomonas dominate soil environments, producing antibiotics, enzymes, and other bioactive compounds as part of ecological competition. Neighboring bacteria counter these with resistance mechanisms such as efflux pumps, enzymatic deactivation, and target modification, forming a baseline resistome predating human antibiotic use.
Genetic Exchange
Plasmids, transposons, and integrons enable horizontal gene transfer (HGT) among soil bacteria, amplifying ARG diversity and persistence across pristine, agricultural, and contaminated soils. Metagenomic studies reveal hundreds of ARG subtypes per soil sample, with multidrug efflux genes often most abundant, varying by ecosystem like tundra or prairie.
Research Value
Profiling soil resistomes via metagenomics tracks ARG origins, propagation via mobile genetic elements (MGEs), and risks of transfer to pathogens, informing One Health strategies to curb antimicrobial resistance spread. Grass phyllospheres may even exceed soil in ARG richness, highlighting interconnected reservoirs.
Aquatic Microbiomes and Gene Flow
Aquatic microbiomes in rivers, lakes, oceans, and sediments form key reservoirs of antibiotic resistance genes (ARGs), mirroring the dynamics seen in soil but amplified by fluid gene flow.
ARG Diversity
These environments host diverse ARGs against β-lactams, tetracyclines, sulfonamides, and more, often in Proteobacteria, Actinobacteria, and abundant taxa like Pelagibacter. Genes persist via plasmids and transposons, with metagenomic surveys revealing target modification (e.g., mutations) in streamlined oligotrophs and efflux pumps elsewhere.
Gene Flow Highways
High water mobility boosts horizontal gene transfer (HGT) through conjugation and transformation, crossing species barriers more readily than in soil. Aquaculture runoff and wastewater enrich ARGs like sul1 and tetA, linking aquatic resistomes to clinical ones.
Influencing Factors
Nutrient loads, pH shifts, salinity, and natural antimicrobials (e.g., from algae) select for specific ARGs; low antibiotic levels still drive HGT and persistence. Studies in Caspian Sea and global oceans underscore their role in microbial evolution and One Health risks.
Applications in Biotechnology
The study of environmental resistomes is not limited to ecology; it has direct implications for biotechnology. Resistance genes discovered in soil and water bacteria can serve as:
Genetic markers for selecting engineered microbial strains
Biosensors to detect specific environmental chemicals
Tools for synthetic biology, enabling controlled gene expression in engineered microbes
By tapping into these hidden genetic libraries, researchers can design microbial systems for bioproduction, bioengineering, and environmental monitoring, turning natural adaptation strategies into practical biotechnological tools.
Bacterial Adaptation and Evolution
Bacterial adaptation in the environmental resistome showcases dynamic evolution through mutation, recombination, and horizontal gene transfer (HGT), enabling rapid responses to stressors like natural antibiotics.
Evolutionary Mechanisms
Resistance genes evolve via point mutations altering targets (e.g., ribosomal modifications), gene duplication creating novel variants, and HGT via plasmids, transposons, integrons, and phages, which shuffle ARGs across species. Biofilm lifestyles versus planktonic growth yield distinct pathways e.g., efflux pump upregulation in biofilms versus target mutations in free cells highlighting lifestyle-driven selection.
Ecological Drivers
Ecological pressures from antibiotic producers (e.g., Streptomyces in soil) and competitors select resilient populations, with microbial density amplifying gene flux in hotspots like soil or aquatic sediments. Community interactions, including exotoxins inducing tolerance states, further steer trajectories toward low-cost, stepwise resistance.
Implications
This fluidity sustains microbial diversity, fostering innovation like multidrug resistance cassettes, but risks pathogen acquisition via HGT, underscoring the resistome's role in global AMR evolution.
Conclusion
The environmental resistome represents a hidden treasure trove of genetic potential in soil and water microbiomes. By studying how resistance genes propagate and function in natural ecosystems, scientists gain insights into microbial adaptation, ecological interactions, and biotechnological applications. Harnessing this genetic diversity opens new avenues for synthetic biology, bioproduction, and environmental biotechnology, transforming naturally occurring bacterial strategies into innovative scientific solutions.
Recent Posts
-
The Intestinal Microbiota: A Complex Microbial Ecosystem
The Intestinal Microbiota: A Complex Microbial Ecosystem The intestinal microbiota represents one of …13th Mar 2026 -
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds
Metagenomics in Environmental Microbiology: Exploring Hidden Microbial Worlds Introduction Microorga …13th Mar 2026 -
The Environmental Resistome: Hidden Genetic Libraries in Soil and Water Microbiomes
The Environmental Resistome : Hidden Genetic Libraries in Soil and Water Microbiomes Introduction Ba …13th Mar 2026