Antibiotic Resistance in Bacteria: A Molecular Evolution Challenge in Modern Microbiology
Antibiotic Resistance in Bacteria: A Molecular Evolution Challenge in Modern Microbiology
Bacteria are among the most adaptable organisms on Earth. Their ability to rapidly evolve allows them to survive in highly competitive environments such as soil ecosystems, marine habitats, industrial bioreactors, and laboratory cultures. One of the most fascinating examples of microbial adaptation is antibiotic resistance.
From a microbiological perspective, antibiotic resistance is not only a challenge but also an extraordinary model of molecular evolution, genetic exchange, and cellular defense mechanisms.
Understanding how bacteria acquire and regulate resistance provides valuable insights into microbial genetics, biotechnology, and environmental microbiology.
What is Antibiotic Resistance?
Antibiotic resistance describes the ability of bacterial cells to survive and grow in the presence of antimicrobial compounds produced by other microorganisms or synthetic molecules.
In natural ecosystems, antibiotics often function as chemical signaling molecules or competitive agents between microbial populations rather than simply as lethal compounds. As a result, bacteria have evolved multiple strategies to neutralize or bypass these molecules.
This phenomenon illustrates the remarkable plasticity of bacterial genomes and their capacity to rapidly adapt to environmental stress.
Figure: WHO bacterial priority pathogens list, 2024 (Organization, W.H, 2024)
Antibiotics as Ecological Molecules
Many antibiotics are naturally produced by microorganisms living in complex microbial communities.
For example, species belonging to the genus Streptomyces synthesize a wide variety of antimicrobial compounds that influence microbial competition in soil ecosystems.
In these environments, resistance genes act as protective mechanisms that allow producing organisms or neighboring microbes to tolerate these compounds.
This ecological perspective highlights that antibiotic resistance is an ancient evolutionary phenomenon, long predating modern biotechnology.
Updates : Click here to read article related
Molecular Mechanisms of Bacterial Resistance
Bacteria employ several sophisticated molecular strategies to resist antimicrobial compounds.
Enzymatic Inactivation
Some bacteria produce enzymes that chemically modify or degrade antimicrobial molecules before they reach their targets.
A well-known example is the production of β-lactamase enzymes, which break down β-lactam compounds synthesized by microorganisms such as Penicillium.
Efflux Pumps
Many bacterial cells contain membrane proteins capable of actively transporting toxic compounds out of the cell.
These efflux pumps reduce intracellular concentrations of antimicrobial molecules and can confer resistance to multiple chemical structures.
Efflux systems are widely studied in environmental microbiology because of their role in cellular detoxification and metabolic regulation.
Target Site Modification
Bacteria may also alter the molecular structures that antimicrobial compounds normally interact with.
Small genetic changes in ribosomal proteins, metabolic enzymes, or membrane structures can significantly reduce the binding efficiency of antimicrobial molecules.
This mechanism demonstrates how minor genetic variations can produce major phenotypic changes in microbial populations.
Figure:Plasmid curing to resensitize the resistant bacteria.
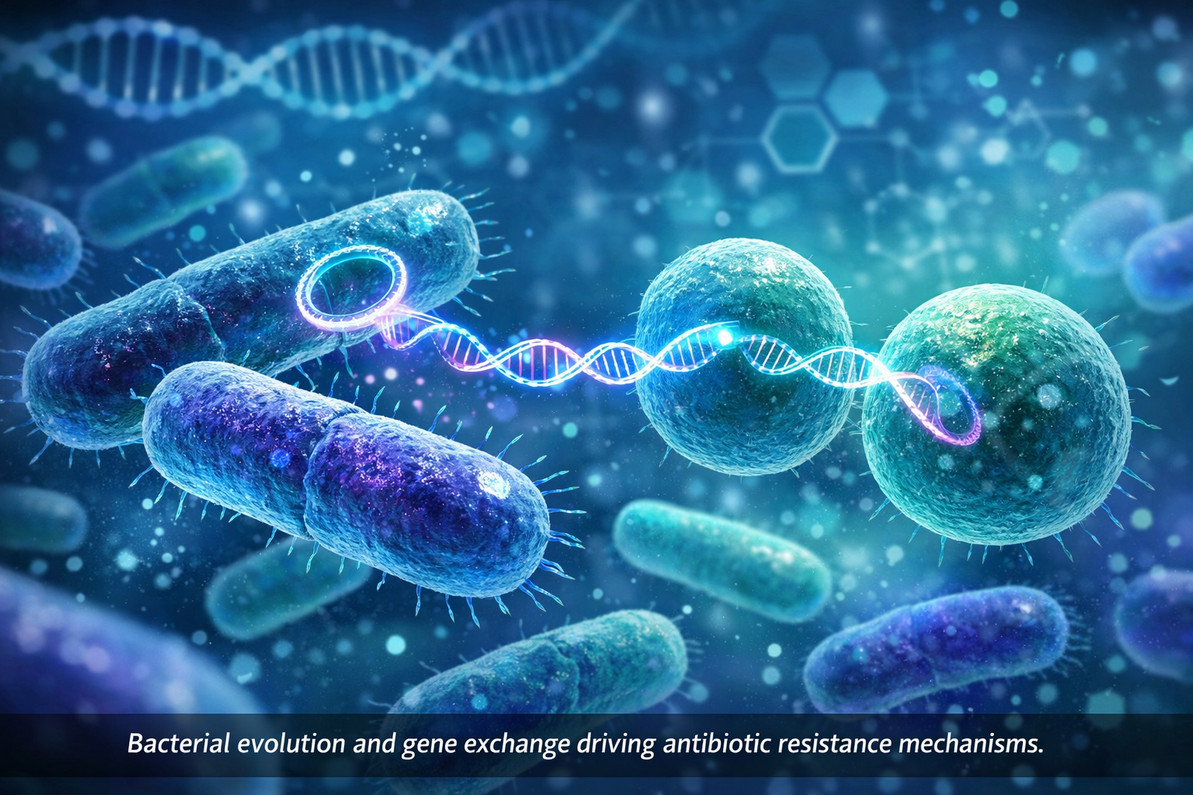
Horizontal Gene Transfer: A Powerful Evolutionary Tool
One of the most remarkable features of bacterial evolution is the ability to exchange genetic information between unrelated cells.
This process, known as Horizontal gene transfer, allows resistance genes to move rapidly through microbial communities.
Three main mechanisms enable this exchange:
Conjugation – transfer of plasmids through direct cell-to-cell contact
Transformation – uptake of extracellular DNA fragments
Transduction – gene transfer mediated by bacteriophages
These processes create highly dynamic microbial populations capable of rapid genetic innovation.
Environmental Reservoirs of Resistance Genes
Natural ecosystems serve as vast reservoirs of resistance genes, sometimes referred to as the environmental resistome.
Soil microbiomes, aquatic systems, and microbial biofilms contain enormous genetic diversity. Many resistance genes discovered in laboratory strains were originally identified in environmental microorganisms.
Studying these reservoirs helps researchers understand how resistance evolves, spreads, and integrates into microbial genomes.
Biotechnological Applications of Resistance Systems
Interestingly, antibiotic resistance genes have become essential tools in molecular biology and biotechnology.
In genetic engineering, resistance markers are frequently used to select successfully transformed cells during cloning or expression experiments.
These markers play a central role in techniques involving organisms such as Escherichia coli, one of the most widely used model organisms in biotechnology laboratories.
Thus, resistance genes are not only evolutionary adaptations but also valuable molecular tools in research and synthetic biology.
Emerging Research Directions
Recent advances in genomics and proteomics are transforming the study of antibiotic resistance.
High-throughput sequencing allows scientists to explore resistance gene diversity across entire microbial communities, while advanced proteomic techniques reveal how resistance proteins function at the molecular level.
Additionally, computational biology is helping researchers map complex networks of resistance genes and regulatory pathways within bacterial cells.
These multidisciplinary approaches are expanding our understanding of microbial adaptation and environmental genetics.
Conclusion
Antibiotic resistance represents a remarkable example of bacterial adaptability and genetic innovation. Rather than being solely a clinical issue, it is fundamentally a microbiological and evolutionary phenomenon rooted in microbial ecology.
By studying resistance mechanisms, scientists gain deeper insights into genome plasticity, molecular defense systems, and microbial community dynamics.
As research tools continue to evolve, antibiotic resistance will remain a powerful model for understanding how microorganisms respond to environmental challenges and shape the microbial world around us.
Recent Posts
-
Advances in Peptide Synthesis Technologies and Their Applications in Modern Biotechnology
Advances in Peptide Synthesis Technologies and Their Applications in Modern Biotechnology …27th Mar 2026 -
Western Blot
Introduction Western blotting is a fundamental analytical technique widely used in molecular biology …27th Mar 2026 -
Ethidium Bromide as a Cooperative Effector of DNA Structure
Introduction DNA structure is not static. Under specific physicochemical conditions, DNA molecules c …25th Mar 2026